Local testing procedures for the functional analysis of variance
Source:R/functional-anova-test.R
functional_anova_test.RdImplements local testing procedures for comparing the mean functions of multiple functional populations in a one-way or multi-way functional analysis of variance (FANOVA) framework. Functional data are tested locally and unadjusted and adjusted p-value functions are provided. The unadjusted p-value function controls the point-wise error rate. The adjusted p-value function can be computed according to the following methods:
global testing (controlling the FWER weakly)
interval-wise testing (controlling the interval-wise error rate)
threshold-wise testing (controlling the FWER asymptotically)
Arguments
- formula
An object of class
stats::formula(or one that can be coerced to that class) specifying the model to be fitted in a symbolic fashion. The response (left-hand side) can be either a matrix of dimension \(n \times J\) containing the pointwise evaluations of \(n\) functions on the same grid of \(J\) points, or an object of classfda::fd.- correction
A string specifying the method used to calculate the adjusted p-value function. Choices are
"Global"for global testing,"IWT"for interval-wise testing, or"TWT"for threshold-wise testing.- dx
A numeric value specifying the discretization step of the grid used to evaluate functional data when it is provided as objects of class
fda::fd. Defaults toNULL, in which case a default value of0.01is used which corresponds to a grid of size100L. Unused if functional data is provided in the form of matrices.- B
An integer value specifying the number of permutations used to evaluate the p-values of the permutation tests. Defaults to
1000L. Passed asn_perminiwt_aov(),twt_aov()andglobal_aov().- method
A string specifying the permutation scheme.
"residuals"permutes residuals under the reduced model (Freedman-Lane scheme);"responses"permutes the responses (Manly scheme). Defaults to"residuals".- recycle
A boolean value specifying whether the recycled version of the interval-wise testing procedure should be used. See Pini and Vantini (2017) for details. Defaults to
TRUE.- stat
A string specifying the test statistic used for the global test.
"Integral"uses the integral of the F-statistic over the domain;"Max"uses the maximum. Defaults to"Integral".
Value
An object of class faov containing the following components:
call: The matched call.design_matrix: The design matrix of the functional ANOVA model.unadjusted_pval_F: A numeric vector of length \(J\) containing the unadjusted p-value function of the global F-test evaluated on the grid.adjusted_pval_F: A numeric vector of length \(J\) containing the adjusted p-value function of the global F-test evaluated on the grid.unadjusted_pval_factors: A numeric matrix with one row per factor containing the unadjusted p-value functions of the per-factor F-tests.adjusted_pval_factors: A numeric matrix with one row per factor containing the adjusted p-value functions of the per-factor F-tests.data_eval: A numeric matrix containing the functional data evaluated on the grid.coeff_regr_eval: A numeric matrix containing the functional regression coefficients evaluated on the grid.fitted_eval: A numeric matrix containing the fitted values of the functional regression evaluated on the grid.residuals_eval: A numeric matrix containing the residuals of the functional regression evaluated on the grid.R2_eval: A numeric vector containing the functional R-squared evaluated on the grid.
Optionally, the list may contain the following components:
pval_matrix_F: A matrix of dimensions \(p \times p\) of p-values of the interval-wise F-tests. Element \((i,j)\) contains the p-value of the test on the interval \((j, j+1, \ldots, j+(p-i))\). Present only ifcorrectionis"IWT".pval_matrix_factors: An array of dimensions \(L \times p \times p\) of p-values of the per-factor interval-wise F-tests. Element \((l,i,j)\) contains the p-value of the joint test on factor \(l\) and interval \((j, j+1, \ldots, j+(p-i))\). Present only ifcorrectionis"IWT".global_pval_F: Global p-value of the overall F-test. Present only ifcorrectionis"Global".global_pval_factors: A numeric vector of global p-values of the per-factor F-tests. Present only ifcorrectionis"Global".
References
Abramowicz, K., Pini, A., Schelin, L., Stamm, A., & Vantini, S. (2022). “Domain selection and familywise error rate for functional data: A unified framework. Biometrics 79(2), 1119-1132.
D. Freedman and D. Lane (1983). A Nonstochastic Interpretation of Reported Significance Levels. Journal of Business & Economic Statistics 1.4, 292-298.
B. F. J. Manly (2006). Randomization, Bootstrap and Monte Carlo Methods in Biology. Vol. 70. CRC Press.
Pini, A., & Vantini, S. (2017). Interval-wise testing for functional data. Journal of Nonparametric Statistics, 29(2), 407-424.
Pini, A., Vantini, S., Colosimo, B. M., & Grasso, M. (2018). Domain‐selective functional analysis of variance for supervised statistical profile monitoring of signal data. Journal of the Royal Statistical Society: Series C (Applied Statistics) 67(1), 55-81.
Abramowicz, K., Hager, C. K., Pini, A., Schelin, L., Sjostedt de Luna, S., & Vantini, S. (2018). Nonparametric inference for functional‐on‐scalar linear models applied to knee kinematic hop data after injury of the anterior cruciate ligament. Scandinavian Journal of Statistics 45(4), 1036-1061.
See also
iwt_aov(), twt_aov() and global_aov() for calling a
specific correction directly. plot.faov() for plotting the results and
summary.faov() for summarizing the results.
Examples
temperature <- rbind(NASAtemp$milan[, 1:100], NASAtemp$paris[, 1:100])
groups <- c(rep(0, 22), rep(1, 22))
# Performing the TWT for multiple populations
TWT_result <- functional_anova_test(
temperature ~ groups,
correction = "TWT",
B = 10L
)
#>
#> ── Point-wise tests ────────────────────────────────────────────────────────────
#>
#> ── Threshold-wise tests ────────────────────────────────────────────────────────
#>
#> ── Threshold-Wise Testing completed ────────────────────────────────────────────
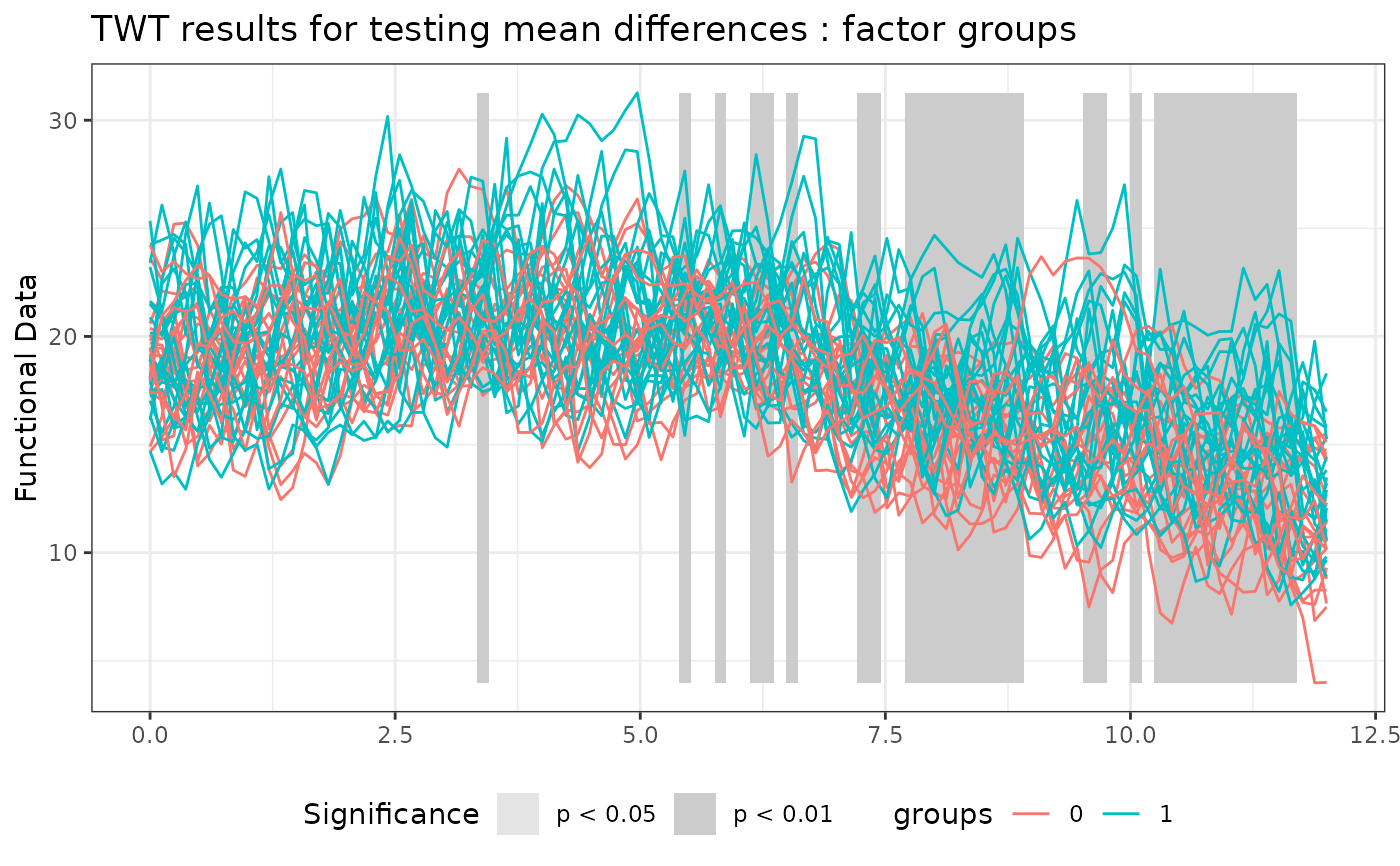
# Plotting the results of the TWT
plot(
TWT_result,
xrange = c(0, 12),
main = "TWT results for testing mean differences"
)
 # Selecting the significant components at 5% level
which(TWT_result$adjusted_pval_F < 0.05)
#> [1] 29 30 40 46 47 48 49 50 55 57 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75
#> [26] 76 77 80 81 84 85 86 87 88 89 90 91 92 93 94 95 96 97
# Performing the IWT for multiple populations
IWT_result <- functional_anova_test(
temperature ~ groups,
correction = "IWT",
B = 10L
)
#>
#> ── Point-wise tests ────────────────────────────────────────────────────────────
#>
#> ── Interval-wise tests ─────────────────────────────────────────────────────────
#>
#> ── Interval-Wise Testing completed ─────────────────────────────────────────────
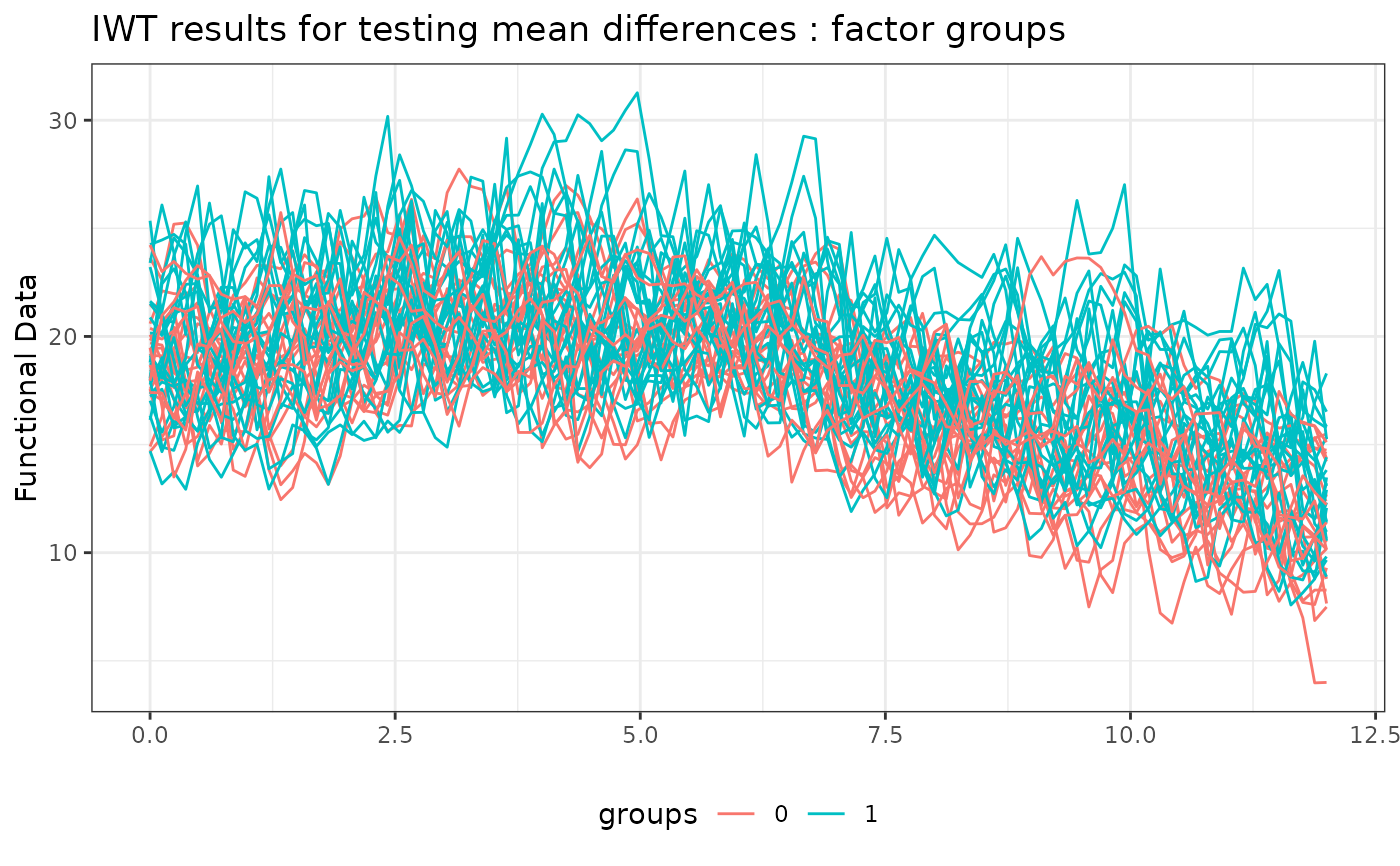
# Plotting the results of the IWT
plot(
IWT_result,
xrange = c(0, 12),
main = "IWT results for testing mean differences"
)
# Selecting the significant components at 5% level
which(TWT_result$adjusted_pval_F < 0.05)
#> [1] 29 30 40 46 47 48 49 50 55 57 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75
#> [26] 76 77 80 81 84 85 86 87 88 89 90 91 92 93 94 95 96 97
# Performing the IWT for multiple populations
IWT_result <- functional_anova_test(
temperature ~ groups,
correction = "IWT",
B = 10L
)
#>
#> ── Point-wise tests ────────────────────────────────────────────────────────────
#>
#> ── Interval-wise tests ─────────────────────────────────────────────────────────
#>
#> ── Interval-Wise Testing completed ─────────────────────────────────────────────
# Plotting the results of the IWT
plot(
IWT_result,
xrange = c(0, 12),
main = "IWT results for testing mean differences"
)
 # Selecting the significant components at 5% level
which(IWT_result$adjusted_pval_F < 0.05)
#> [1] 69 70 71 72 88 89 90
# Selecting the significant components at 5% level
which(IWT_result$adjusted_pval_F < 0.05)
#> [1] 69 70 71 72 88 89 90