One population Interval Wise Testing procedure
Source:R/ITP1bspline.R, R/ITP1fourier.R, R/os-iwt.R
IWT1.RdThe function implements the Interval Wise Testing procedure for testing the center of symmetry of a functional population. Functional data are tested locally and unadjusted and adjusted p-value functions are provided. The unadjusted p-value function controls the point-wise error rate. The adjusted p-value function controls the interval-wise error rate.
Usage
ITP1bspline(data, mu = 0, B = 1000, order = 2, nknots = dim(data)[2])
ITP1fourier(data, mu = 0, B = 1000, maxfrequency = floor(dim(data)[2]/2))
iwt1(
data,
mu = 0,
n_perm = 1000L,
dx = NULL,
recycle = TRUE,
verbose = FALSE,
aggregation_strategy = c("integral", "max")
)
IWT1(data, mu = 0, B = 1000L, dx = NULL, recycle = TRUE)Arguments
- data
Either a numeric matrix or an object of class
fda::fdspecifying the sample data. If the data is provided within a matrix, it should be of shape \(n \times J\) and it should contain in each row one of the \(n\) functions in the sample and in columns the evaluation of each function on a same uniform grid of size \(J\).- mu
Either a numeric value or a numeric vector or an object of class
fda::fdspecifying the functional center of symmetry under the null hypothesis. Ifmuis a constant, then a constant function is used. Ifmuis a numeric vector, it must correspond to evaluations of the center function on the same grid that has been used to evaluate the data sample. Defaults to0.- B
The number of iterations of the MC algorithm to evaluate the p-values of the permutation tests. Defaults to
1000L.- order
Order of the B-spline basis expansion. Defaults to
2L.- nknots
Number of knots of the B-spline basis expansion. Defaults to
dim(data)[2].- maxfrequency
The maximum frequency to be used in the Fourier basis expansion of data. Defaults to
floor(dim(data)[2] / 2), leading to an interpolating expansion.- n_perm
An integer value specifying the number of permutations to use for the local testing procedure. Defaults to
1000L.- dx
A numeric value specifying the step of the uniform grid on which the data are evaluated. If
NULL, the step is automatically inferred from the data. Defaults toNULL.- recycle
A boolean value specifying whether to recycle the test statistic values across permutations for the IWT procedure. Defaults to
TRUE.- verbose
A boolean value specifying whether to print the progress of the computation. Defaults to
FALSE.- aggregation_strategy
A string specifying the strategy to aggregate the point-wise test statistics for the correction procedure. Possible values are
"integral"and"max". Defaults to"integral".
Value
An object of class fos containing the following components:
data: A numeric matrix of shape \(n \times J\) containing the evaluation of the \(n\) functions on a uniform grid of size \(J\).mu: A numeric vector of shape \(J\) containing the evaluation of the functional center of symmetry under the null hypothesis on the same uniform grid used to evaluate the functional sample.unadjusted_pvalues: A numeric vector of size \(J\) containing the evaluation of the unadjusted p-value function on the same uniform grid used to evaluate the functional sample.adjusted_pvalues: A numeric vector of size \(J\) containing the evaluation of the adjusted p-value function on the same uniform grid used to evaluate the functional sample.correction_method: A string containing the correction method used to compute the adjusted p-value function.
Optionally, the list may contain the following component:
pvalue_matrix: A numeric matrix of shape \(p \times p\) containing the p-values of the interval-wise tests. Element \(i, j\) contains the p-value of the test performed on the interval indexed by \(j, j+1, \dots, j+(p-i)\). Only present if thecorrectionargument is set to"IWT".
References
Pini, Alessia, and Simone Vantini. 2016. "The interval testing procedure: a general framework for inference in functional data analysis." Biometrics 72 (3): 835–845.
Pini, Alessia, and Simone Vantini. 2017. "Interval-Wise Testing for Functional Data." Journal of Nonparametric Statistics 29 (2): 407–24.
See also
functional_one_sample_test() for the interface function and
plot.fos() and IWTimage() for plotting the results.
Examples
# Performing the IWT for one population
IWT_result <- iwt1(NASAtemp$paris, mu = 4, n_perm = 10L)
# Selecting the significant components at 5% level
which(IWT_result$adjusted_pvalues < 0.05)
#> [1] 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18
#> [19] 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36
#> [37] 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54
#> [55] 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72
#> [73] 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90
#> [91] 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108
#> [109] 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126
#> [127] 127 128 129 130 131 132 133 134 135 136 137 138 261 262 263 264 265 266
#> [145] 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284
#> [163] 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302
#> [181] 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320
#> [199] 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338
#> [217] 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356
#> [235] 357 358 359 360 361 362 363 364 365
# Performing the IWT for one population
IWT_result <- IWT1(NASAtemp$paris, mu = 4, B = 10L)
# Plotting the results of the IWT
plot(IWT_result, xrange = c(0, 12), main = "Paris temperatures")
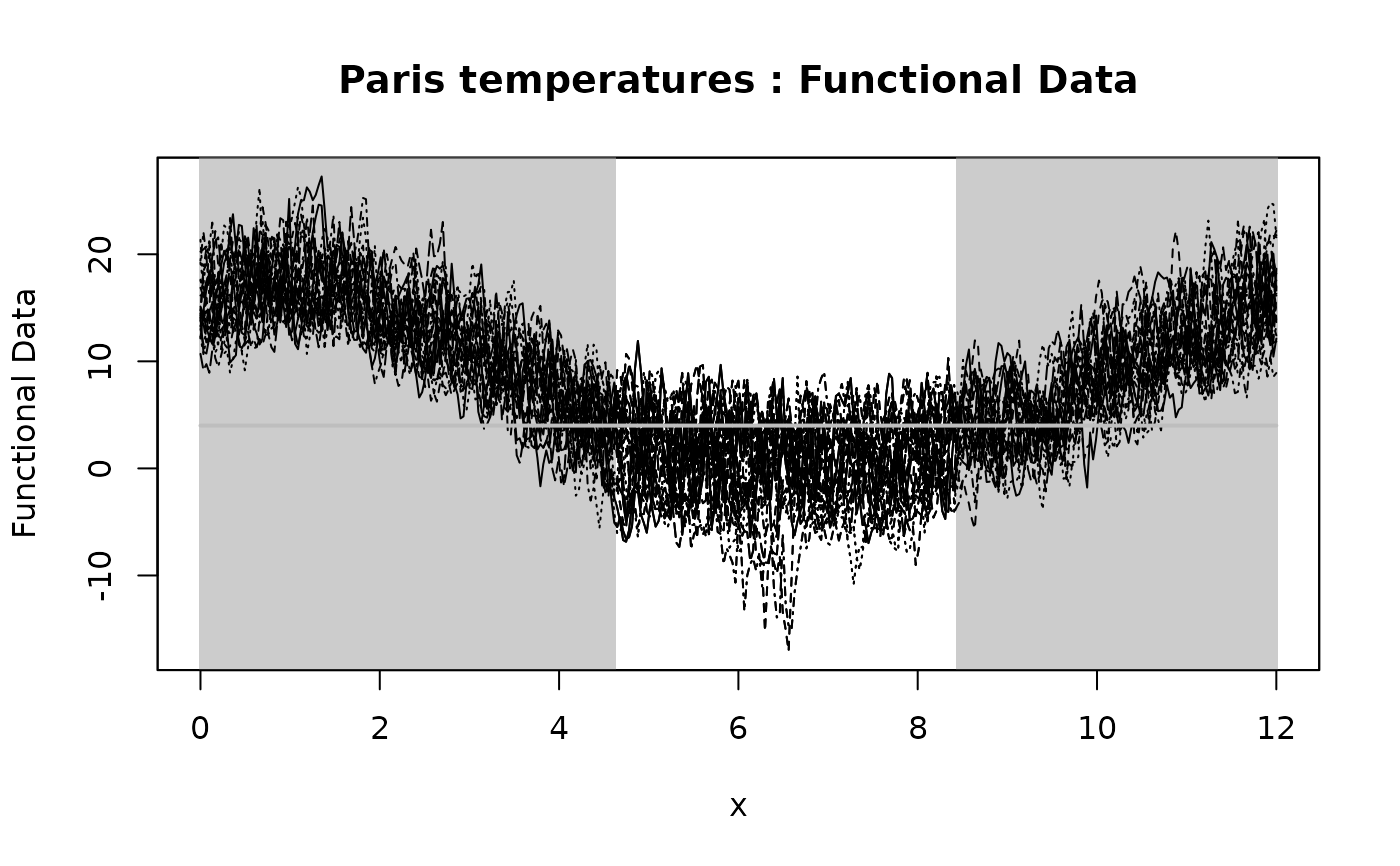 # Plotting the p-value heatmap
IWTimage(IWT_result, abscissa_range = c(0, 12))
# Plotting the p-value heatmap
IWTimage(IWT_result, abscissa_range = c(0, 12))
 # Selecting the significant components at 5% level
which(IWT_result$adjusted_pval < 0.05)
#> [1] 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18
#> [19] 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36
#> [37] 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54
#> [55] 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72
#> [73] 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90
#> [91] 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108
#> [109] 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126
#> [127] 127 128 129 130 131 132 133 134 135 136 258 259 260 261 262 263 264 265
#> [145] 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283
#> [163] 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301
#> [181] 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319
#> [199] 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337
#> [217] 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355
#> [235] 356 357 358 359 360 361 362 363 364 365
# Selecting the significant components at 5% level
which(IWT_result$adjusted_pval < 0.05)
#> [1] 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18
#> [19] 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36
#> [37] 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54
#> [55] 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72
#> [73] 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90
#> [91] 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108
#> [109] 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126
#> [127] 127 128 129 130 131 132 133 134 135 136 258 259 260 261 262 263 264 265
#> [145] 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283
#> [163] 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301
#> [181] 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319
#> [199] 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337
#> [217] 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355
#> [235] 356 357 358 359 360 361 362 363 364 365